Federated Learning for Sparse Bayesian Models with Applications to Electronic Health Records and Genomics
Abstract
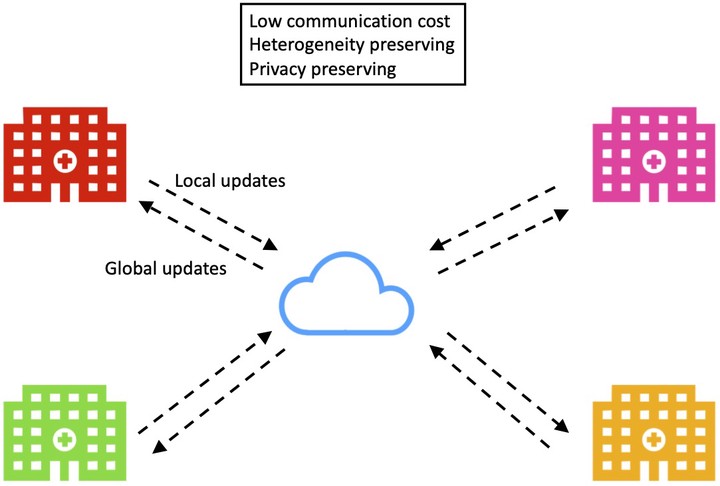
Federated learning is becoming increasingly more popular as the concern of privacy breaches rises across disciplines including the biological and biomedical fields. The main idea is to train models locally on each server using data that are only available to that server and aggregate the model (not data) information at the global level. While federated learning has made significant advancements for machine learning methods such as deep neural networks, to the best of our knowledge, its development in sparse Bayesian models is still lacking. Sparse Bayesian models are highly interpretable with natural uncertain quantification, a desirable property for many scientific problems. However, without a federated learning algorithm, their applicability to sensitive biological/biomedical data from multiple sources is limited. Therefore, to fill this gap in the literature, we propose a new Bayesian federated learning framework that is capable of pooling information from different data sources without breaching privacy. The proposed method is conceptually simple to understand and implement, accommodates sampling heterogeneity (i.e., non-iid observations) across data sources, and allows for principled uncertainty quantification. We illustrate the proposed framework with three concrete sparse Bayesian models, namely, sparse regression, Markov random field, and directed graphical models. The application of these three models is demonstrated through three real data examples including a multi-hospital COVID-19 study, breast cancer protein-protein interaction networks, and gene regulatory networks.